Note
Click here to download the full example code
GroupLasso for linear regression with dummy variables¶
A sample script for group lasso with dummy variables
Setup¶
import matplotlib.pyplot as plt
import numpy as np
from sklearn.linear_model import Ridge
from sklearn.metrics import r2_score
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import OneHotEncoder
from group_lasso import GroupLasso
from group_lasso.utils import extract_ohe_groups
np.random.seed(42)
GroupLasso.LOG_LOSSES = True
Set dataset parameters¶
num_categories = 30
min_options = 2
max_options = 10
num_datapoints = 10000
noise_std = 1
Generate data matrix¶
X_cat = np.empty((num_datapoints, num_categories))
for i in range(num_categories):
X_cat[:, i] = np.random.randint(min_options, max_options, num_datapoints)
ohe = OneHotEncoder()
X = ohe.fit_transform(X_cat)
groups = extract_ohe_groups(ohe)
group_sizes = [np.sum(groups == g) for g in np.unique(groups)]
active_groups = [np.random.randint(0, 2) for _ in np.unique(groups)]
Generate coefficients¶
w = np.concatenate(
[
np.random.standard_normal(group_size) * is_active
for group_size, is_active in zip(group_sizes, active_groups)
]
)
w = w.reshape(-1, 1)
true_coefficient_mask = w != 0
intercept = 2
Generate regression targets¶
y_true = X @ w + intercept
y = y_true + np.random.randn(*y_true.shape) * noise_std
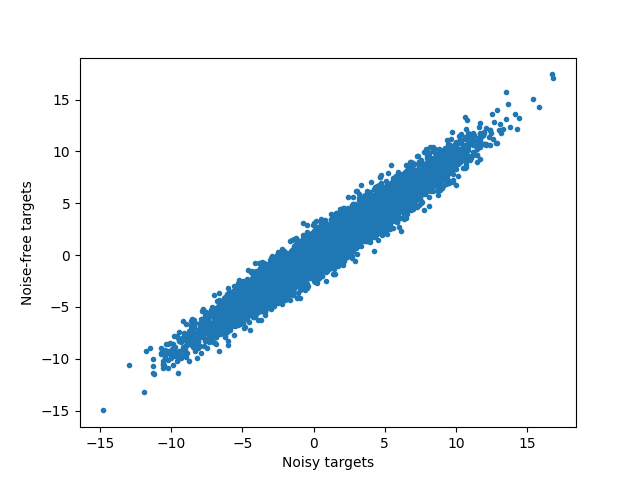
View noisy data and compute maximum R^2¶
plt.figure()
plt.plot(y, y_true, ".")
plt.xlabel("Noisy targets")
plt.ylabel("Noise-free targets")
# Use noisy y as true because that is what we would have access
# to in a real-life setting.
R2_best = r2_score(y, y_true)
Generate pipeline and train it¶
pipe = pipe = Pipeline(
memory=None,
steps=[
(
"variable_selection",
GroupLasso(
groups=groups,
group_reg=0.1,
l1_reg=0,
scale_reg=None,
supress_warning=True,
n_iter=100000,
frobenius_lipschitz=False,
),
),
("regressor", Ridge(alpha=1)),
],
)
pipe.fit(X, y)
Out:
Pipeline(steps=[('variable_selection',
GroupLasso(group_reg=0.1,
groups=array([ 0., 0., 0., 0., 0., 0., 0., 0., 1., 1., 1., 1., 1.,
1., 1., 1., 2., 2., 2., 2., 2., 2., 2., 2., 3., 3.,
3., 3., 3., 3., 3., 3., 4., 4., 4., 4., 4., 4., 4.,
4., 5., 5., 5., 5., 5., 5., 5., 5., 6., 6., 6., 6.,
6., 6., 6., 6., 7., 7., 7., 7., 7., 7., 7., 7., 8.,
8., 8., 8., 8., 8., 8., 8., 9., 9., 9., 9., 9., 9.,
9., 9., 10., 10., 10., 10., 10., 10., 10., 10., 1...
21., 21., 21., 21., 21., 21., 21., 22., 22., 22., 22., 22., 22.,
22., 22., 23., 23., 23., 23., 23., 23., 23., 23., 24., 24., 24.,
24., 24., 24., 24., 24., 25., 25., 25., 25., 25., 25., 25., 25.,
26., 26., 26., 26., 26., 26., 26., 26., 27., 27., 27., 27., 27.,
27., 27., 27., 28., 28., 28., 28., 28., 28., 28., 28., 29., 29.,
29., 29., 29., 29., 29., 29.]),
l1_reg=0, n_iter=100000, scale_reg=None,
supress_warning=True)),
('regressor', Ridge(alpha=1))])
Extract results and compute performance metrics¶
# Extract from pipeline
yhat = pipe.predict(X)
sparsity_mask = pipe["variable_selection"].sparsity_mask_
coef = pipe["regressor"].coef_.T
# Construct full coefficient vector
w_hat = np.zeros_like(w)
w_hat[sparsity_mask] = coef
R2 = r2_score(y, yhat)
# Print performance metrics
print(f"Number variables: {len(sparsity_mask)}")
print(f"Number of chosen variables: {sparsity_mask.sum()}")
print(f"R^2: {R2}, best possible R^2 = {R2_best}")
Out:
Number variables: 240
Number of chosen variables: 144
R^2: 0.9278329616431523, best possible R^2 = 0.9394648554757948
Visualise regression coefficients¶
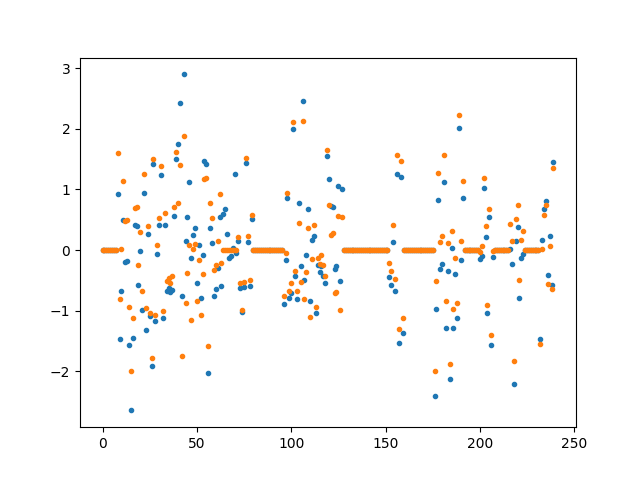
for i in range(w.shape[1]):
plt.figure()
plt.plot(w[:, i], ".", label="True weights")
plt.plot(w_hat[:, i], ".", label="Estimated weights")
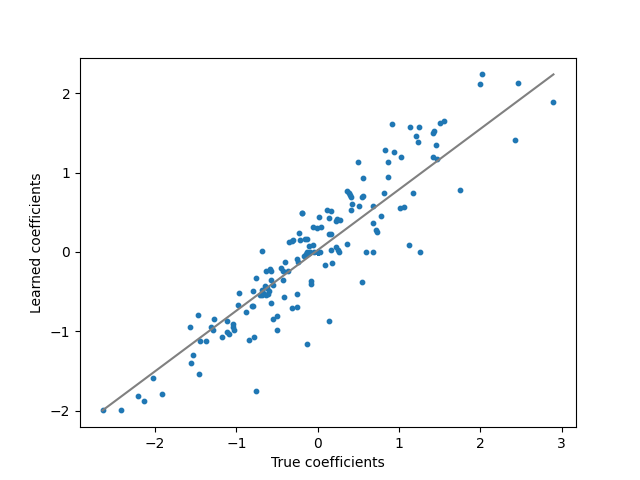
plt.figure()
plt.plot([w.min(), w.max()], [coef.min(), coef.max()], "gray")
plt.scatter(w, w_hat, s=10)
plt.ylabel("Learned coefficients")
plt.xlabel("True coefficients")
plt.show()
Total running time of the script: ( 0 minutes 18.205 seconds)